Human genetic variation in APOE (Apolipoprotein E) and MAPT gene (Microtubule Associated Protein Tau) significantly influences risk for Alzheimer’s disease (AD) and tauopathies including frontotemporal dementia (FTD) and progressive supranuclear palsy. APOE alleles (ε2, ε3, ε4) primarily affect glial support and amyloid handling, while MAPT haplotypes (H1 risk, H2 protective in most primary tauopathies) modulate tau isoform balance and expression. These variants drive cell-type-specific phenotypes, with astrocytes supporting neuronal lipid and metabolic needs via APOE, and tau pathology predominantly neuronal but extending to glia. Genetically defined human primary cells, stratified by these genotypes, offer native platforms for modeling sporadic disease, especially as MAPT-genotyped primary neurons and astrocytes become commercially available.
MAPT Genotype/Haplotype Effects
In Neurons: The MAPT locus features H1 (risk for PSP, CBD, some AD/PD cases) and H2 haplotypes, separated by a ~900 kb inversion. H1 elevates total MAPT expression (~1.5-fold) and biases splicing toward 4R tau isoforms (exon 10 inclusion), promoting microtubule instability, tau aggregation into NFTs, and impaired axonal transport. H1 subhaplotypes (e.g., H1c) further increase 4R bias. In contrast, H2 favors 3R isoforms and is protective in most 4R tauopathies but may increase risk for 3R-dominant Pick’s disease. Some iPSC models show subtle or maturation-dependent differences, but postmortem and allele-specific expression data confirm H1-driven 4R excess as a key mechanism for tauopathy vulnerability.
In Astrocytes: Though tau is neuron-enriched, astrocytes express low MAPT levels and accumulate pathological tau in tauopathies. H1 haplotypes associate with altered glial tau mRNA/protein, increased astrocytic proportions in vulnerable brain regions, and transcriptional shifts toward inflammation (e.g., tp53, histone pathways) and chemotaxis dysregulation. Neighboring 17q21.31 genes (e.g., LRRC37A/2, reduced in H1) affect astrocytic function, including pro-inflammatory responses and interaction with α-synuclein in synucleinopathies/PD. H1 may enhance astrocytic tau uptake/seeding and gliosis, contributing to non-cell-autonomous spread and neuroinflammation.
MAPT haplotypes thus extend risk beyond neurons, disrupting glial homeostasis and amplifying pathology.
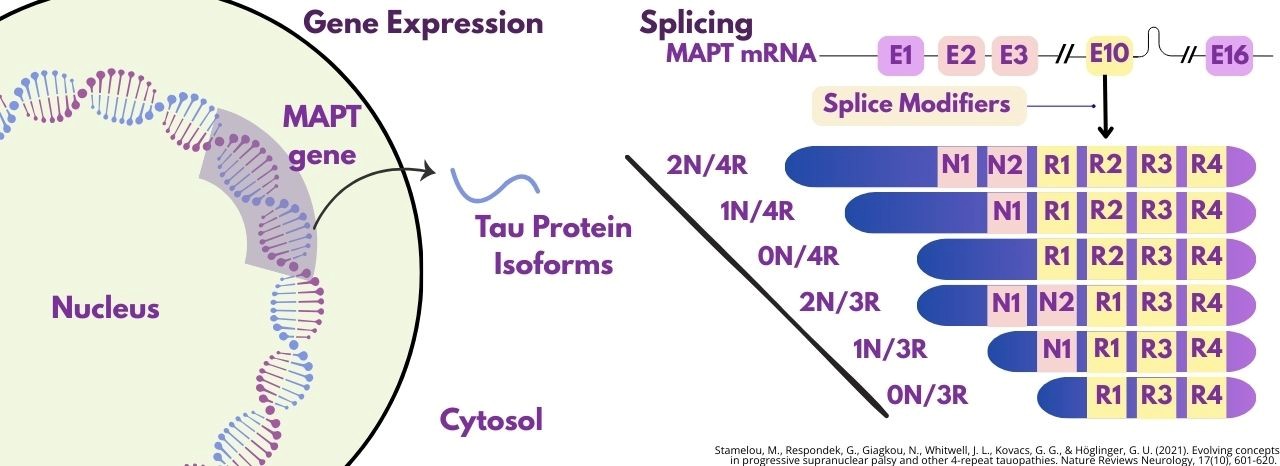
Genetic variants in MAPT affect the expression of tau protein. MAPT haplotypes influence alternative splicing of exon 10, which determines the balance of 3R and 4R tau isoforms (Figure 1).
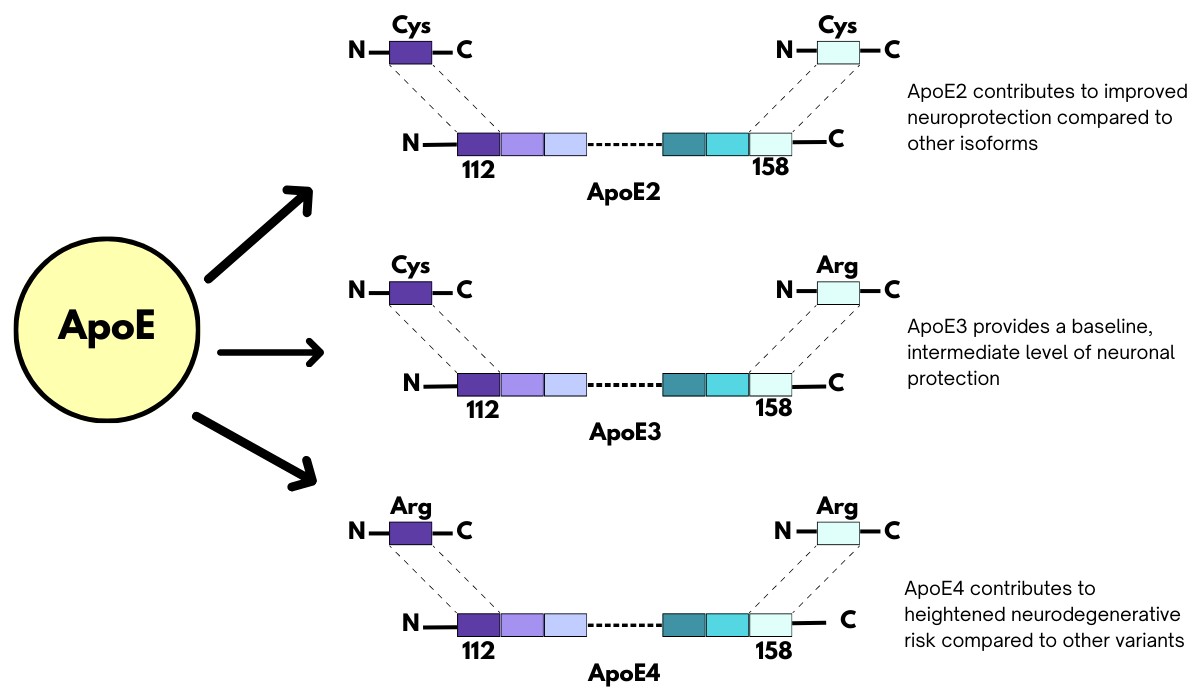
APOE Genotype Effects
In Astrocytes: Astrocytes produce most brain APOE, lipidating particles to supply cholesterol to neurons. The ε4 allele (major AD risk factor) impairs lipidation, reduces cholesterol efflux, and causes intracellular accumulation, downregulating SREBP2 pathways. APOE4 astrocytes show heightened pro-inflammatory signaling (e.g., C3 secretion, A1-like reactivity) and reduced Aβ clearance via LRP1, prolonging amyloid half-life and promoting plaque seeding.
In Neurons: APOE4 indirectly harms neurons through poor astrocytic support, elevating tau hyperphosphorylation (via reduced PP2A or GSK3β dysregulation), decreasing synaptic density and LTP, and disrupting metabolic coupling (e.g., lactate shuttling failure, increased ROS). Direct neuronal APOE4 effects include stress granule accumulation and synaptic proteome alterations. These changes position APOE4 astrocytes as drivers of neurotoxic environments, accelerating AD pathology.
Importance of Genetically Defined Human Primary Neurons and Astrocytes
iPSC-derived models enable isogenic manipulation but often show variability related to reprogramming, clonal selection, loss of native epigenetic context, and extended differentiation timelines. In contrast, human primary neurons and astrocytes retain their native cellular identity without genetic reprogramming, providing a more direct representation of in vivo cell biology.
Primary neurons and astrocytes exhibit stable neuronal or glial lineage commitment, physiologically relevant MAPT expression, and reproducible responses to experimental perturbation. When these cells are genetically defined for APOE or MAPT haplotypes, researchers can stratify experiments by known risk-associated genetic backgrounds while avoiding artifacts introduced by iPSC reprogramming or gene editing.
Genotyping for H1/H2 haplotypes and the H1c sub-haplotype enables controlled comparisons of tau regulation, isoform balance, stress responsiveness, and neuron–astrocyte interactions across defined genetic backgrounds. Even in early developmental contexts, MAPT haplotype–dependent differences can influence tau expression levels and cellular responses, providing insight into genetically driven vulnerability that may later manifest in disease.
Genetically stratified primary neurons and astrocytes support co-culture and mechanistic studies of tau biology, glial modulation, and neuroinflammatory signaling in a human-relevant system. By controlling donor genetics at the outset, researchers can reduce variability, improve reproducibility, and better interpret genotype-dependent effects in neurodegenerative disease research.
At ScienCell’s Research Laboratories, we are excited to now offer human primary neurons and astrocytes genotyped for key neurodegeneration-associated variants, including MAPT haplotypes (H1/H2 and H1c) and APOE genotypes. Researchers can request specific genotypes and select donor lots that match the genetic background required for their studies. Combining physiologically relevant human primary cells with verified donor genotypes, provides an alternative to engineered models, enabling genetically informed experimental design while preserving native cellular biology.
For inquiries, please reach out via email, sales@sciencellonline.com or call us at 1.877.602.8549.
APOE kit also available
We also offer APOE Genotyping qPCR Assay Kit designed to complement APOE genotyping.
References:
- Myers, A. J., Kaleem, M., Marlowe, L., Pittman, A. M., Lees, A. J., Hardy, J., et al. (2005). MAPT haplotypes: At the crossroads of genetic risk and tau expression in the human brain. Human Molecular Genetics, 14(9), 1353–1362. https://doi.org/10.1093/hmg/ddi146
- Pittman, A. M., Myers, A. J., Abou-Sleiman, P., Fung, H. C., Kaleem, M., Marlowe, L., et al. (2005). Linkage disequilibrium fine mapping and haplotype association analysis of the MAPT locus in neurodegenerative disease. Brain, 128(8), 1830–1839. https://doi.org/10.1093/brain/awh561
- Trabzuni, D., Wray, S., Vandrovcova, J., Ramasamy, A., Walker, R., Smith, C., et al. (2012). MAPT expression and splicing is differentially regulated by brain region: Relation to genotype and implication for tauopathies. Brain, 135(7), 2169–2177. https://doi.org/10.1093/brain/aws143
- Liu, C. C., Liu, C. C., Kanekiyo, T., Xu, H., & Bu, G. (2013). Apolipoprotein E and Alzheimer disease: Risk, mechanisms and therapy. Nature Reviews Neurology, 9(2), 106–118. https://doi.org/10.1038/nrneurol.2012.263
- Shi, Y., Yamada, K., Liddelow, S. A., Smith, S. T., Zhao, L., Luo, W, et al. (2017). ApoE4 markedly exacerbates tau-mediated neurodegeneration in a mouse model of tauopathy. Nature, 549(7673), 523–527. https://doi.org/10.1038/nature24016
- Shi, Y., & Holtzman, D. M. (2018). Interplay between innate immunity and Alzheimer disease: APOE and TREM2 in the spotlight. Neuron, 97(1), 28–42. https://doi.org/10.1016/j.neuron.2017.11.026
- Karch, C. M., Cruchaga, C., & Goate, A. M. (2014). Alzheimer’s disease genetics: From the bench to the clinic. Neuron, 83(1), 11–26. https://doi.org/10.1016/j.neuron.2014.05.041
- Lee, S.-I., Jeong, W., Lim, H., Cho, S, Lee, H., Jang, Y., et al. (2021). APOE4-carrying human astrocytes oversupply cholesterol to promote neuronal lipid raft expansion and Aβ generation. Stem Cell Reports, 16(9), 2128–2137. https://doi.org/10.1016/j.stemcr.2021.07.017